ISSN 2410-5708 / e-ISSN 2313-7215
Year 13 | No. 38 | October 2024- January 2025
© Copyright (2024). National Autonomous University of Nicaragua, Managua.
This document is under a Creative Commons
Attribution-NonCommercial-NoDerivs 4.0 International licence.
GelRed dye for staining nucleic acids in agarose gel was used in the molecular biology laboratory
https://doi.org/ 10.5377/rtu.v13i38.19325
Submitted on september 26th, 2024 / Accepted on october 10th, 2024
Oscar Arbizú Medina
Professor UNAN-Managua/ Head of the Molecular Biology
Laboratory "M. Elmer Cisneros in memoriam", Area of Knowledge
of Health Sciences, UNAN-Managua
Francisco Romero Oviedo
Research Professor Laboratory of Molecular Biology "M. Elmer
Cisneros in memoriam", Area of Knowledge of Health Sciences,
UNAN-Managua
Yosimar Narváez González
Researcher Laboratory of Molecular Biology "M. Elmer Cisneros
in memoriam", Area of Knowledge of Molecular Biology Health,
UNAN-Managua
Roshell Rizo Obregón
Researcher Molecular Biology Laboratory "MA. Elmer Cisneros
in memoriam", Polytechnic Institute of Health, National
Autonomous University of Nicaragua, Managua, UNAN-Managua
Section:
Scientific research article
Keywords:
Abstract
Ethidium bromide (EtBr) is a fluorescent dye widely used for staining nucleic acids in agarose and polyacrylamide gels. However, EtBr has several drawbacks, such as its high toxicity and mutagenic capacity, which restricts its use and management. In this study, GelRed®, an alternative dye developed by Biotium, which offers greater sensitivity, safety, and stability than EtBr, was evaluated. The performance of both dyes was compared for the staining of DNA amplified by PCR, using the protocols of BCR-ABL, JACK2, ESBL, carbapenemases IMP, VIM, SPM, GIM, NDM, KPC, OXA, mcr-1/mcr-2 and mecca.
The results indicated that GelRed® can be used to visualize DNA with a fluorescence intensity equal to or greater than that of EtBr, without the need to modify the extraction, amplification, and electrophoresis protocols. Furthermore, GelRed® did not interfere with restriction enzymes and PCR and does not exhibit mutagenic or cytotoxic effects. It is concluded that GelRed® is a safe and sensitive option compared to EtBr for staining nucleic acids in agarose gels.
Introduction
Nucleic acid staining in agarose and polyacrylamide gels is a basic and essential technique in molecular biology laboratories, to visualize DNA or RNA fragments that are separated by electrophoresis and visualized by ultraviolet light.
The most commonly used dye for nucleic acid staining in gels is ethidium bromide (EtBr), a substance that is inserted between the DNA bases of DNA or RNA, and that fluoresces orange-red when exposed to ultraviolet (UV) light. The EtBr is inexpensive, easy to use, and compatible with most imaging systems, giving it an advantage (Hall, 2019).
However, EtBr also has several drawbacks, which limit its use and handling. EtBr is a mutagenic and cytotoxic agent that can cause damage to DNA and living cells, posing a cancer risk to both users and the environment (Ude et al., 2022). Consequently, EtBr must be handled, stored, and disposed of with protective and precautionary measures. Due to these drawbacks, several alternative dyes to EtBr have been developed, which offer greater sensitivity, safety, and stability. Among the different dyes, GelRed® stands out, a dye created by Biotium, which covalently binds to DNA or RNA and produces fluorescence when exposed to UV light (Masciotti, 2018).
GelRed® is impervious to living cells and contains no mutagenic or toxic substances, reducing the risk of exposure to users and the environment. (Debroy et al., 2022) Similarly, it has a higher affinity for DNA and RNA than EtBr, which translates into higher sensitivity and a lower concentration needed for staining, it does not affect restriction enzymes or PCR, so it can be used in samples before or after electrophoresis, without having an impact on subsequent reactions (Haines et al., 2015) (Hall, 2019).
In this study, the stability of the use of GelRed® for DNA staining in agarose gels in the different protocols was evaluated; BCR-ABL, JACK2, BLEE, Carbapenemases IMP, VIM, SPM, GIM, NDM, KPC, OXA, mcr-1/mcr-2, and mecA, which are applied in the Elmer Cisneros in memorial molecular biology laboratory. A comparison was made between the performance of GelRed® and EtBr, considering its sensitivity, reproducibility, and efficiency. Pre-load staining and staining of loaded samples were used as the most common methods for staining nucleic acids in gels. The data and results obtained were analyzed, and the advantages and disadvantages of each dye were discussed.
Materials and methods
DNA extraction.
Blood DNA extraction was performed following the manufacturer’s protocol for DNA extraction using the QIAamp DNA Mini Kit (Qiagen). For bacterial DNA extraction, cultures were used on TSA agar, from which a CFU pool was obtained to later perform the heat extraction method. The extracted DNA was quantified using a Nanodrop Lite 2763 and its quality and integrity were verified (Vílchez & Alonso, 2009).
DNA amplification by PCR.
DNA fragments associated with the genes BCR-ABL, JACK2, BLEE, Carbapenemases IMP, VIM, SPM, GIM, NDM, KPC, OXA, mcr-1/mcr-2 and mecA were amplified, using the primers and reaction conditions specified by Arbizú et al. 2023, and the manufacturer’s indications of the PCR kit GoTaq Green Master Mix (Promega). Amplification was performed in a Thermal Cycler (Fisherbrand), and the size and specificity of PCR products were verified by 1.5% agarose gel electrophoresis (Arbizú Medina et al., 2023).
Agarose gel and nucleic acid staining.
In the preparation of the gel, agarose was used, with a concentration of 1.5% in TBE buffer (Tris-Borate-EDTA), to be used as a base in nucleic acid electrophoresis. Two different methods were used that were adapted to the staining technique. To perform the post-electrophoresis staining with GelRed®, gels were prepared without adding any type of dye, and then mixed the dye with the Loading Buffer in a concentration of 1:9. This mixture was used to load the samples into the agarose gel, using 2ul of Buffer+GelRed® and 6ul of the amplified PCR products. However, to stain a pre-electrophoresis gel, Ethidium Bromide was incorporated into the agarose by pre-staining. This was done by adding 0.5 μg/mL of EtBR to 100 mL of agarose after melting. In both methods, the electrophoresis technique was performed with a condition of 120 volts at 45-60 minutes depending on the protocol used, at the end of the electrophoresis the bands were visualized using an ultraviolet light transluminator and they were photographed with a photodocumenter.
Results
Figure 1 shows the efficiency between the images of the gels stained with EtBr and GelRed® for the laboratory protocol and shows that both dyes allowed the visualization of the amplified PCR products and their base pairs, with a similar fluorescence intensity. In Figure 2, the results indicate that the dye GelRed® has a similar or slightly higher quality than EtBr for DNA staining in agarose gels, unlike EtBr, GelRed®does not have the ability to cross the cell membrane, thus guaranteeing the safety of laboratory personnel who handle this type of chemical avoiding the risk of cellular alterations.
Figure 1 EtBr. Well1. Molecular marker, well 2 positive control, well 3 to 13 patient, 14 negative control
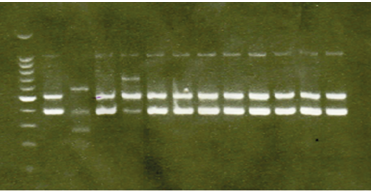
Figure 2 GelRed Well1. Molecular marker, well 2 positive control, well 3 to 10 patients, 11 negative control.
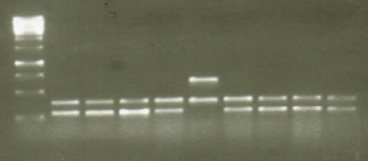
Figure 3. Comparison of gelred reproducibility for DNA staining in agorose gels, DNA amplified by gelred-stained laboratory protocols by the gelred staining method of buffer-loaded samples. The samples correspond. The molecular weight marker is 100bp. He also evaluated the efficiency of both dyes for DNA staining in agarose gels, using the protocols to find bacterial resistance genes respectively. 7 ul of each sample were loaded into the wells of a 1.5% agarose gel. The gel was stained with gelred, following the methods of staining the gel and staining the samples loaded in the buffer. The gel was visualized on a UV transilluminator, and the image was captured with a digital camera.
Discussion
The comparative study between GelRed® and EtBr, fluorescent dyes traditionally used in electrophoresis, through this study it has been shown that GelRed® offers a safer and more efficient alternative since it exhibits a fluorescence intensity comparable to or even higher than that of EtBr, which allows the clear visualization of the DNA or RNA bands in the gels. GelRed® is considered a much safer dye than EtBr, as it has no toxicity or mutagenic effects. In addition, it does not interfere with restriction enzymes or PCR, allowing it to be used in a wide range of applications in electrophoresis.
Unlike the comparative study by (Hall, A. C. (2019), in our study we did not fuse the Gelred® with the agarose, but a solution was prepared with the loading buffer (Gelred® /loading) in a 1:9 ratio, on the other hand, the EtBr had to be fused with the agar to stain the gel and the amplified DNA could be observed, generating a risk due to the evaporation emitted from the EtBr to the users.
Conclusions
In conclusion, the results support the substitution of ethidium bromide (EtBr) by GelRed® in the staining of nucleic acids in agarose gels. GelRed® showed a fluorescence intensity comparable to or greater than EtBr without the need for protocol adjustments, and its inherent safety positions it as a prominent choice. This transition improves safety in the laboratory and optimizes the sensitivity and stability of electrophoresis techniques, contributing significantly to the efficiency of procedures in molecular biology.
Figure 3. Gelred Well 1. Molecular marker, well 2 positive control, well 3 to 12 patient, 11 negative control.

References
Arbizú Medina, O., Romero Oviedo, F., García Rosales, K., Molina Morales, A. E., García Herrera, F. A., Centeno Rizo, B. R., & Amaya, E. (2023). Acinetobacter baumannii is multiresistant to carbapenems at the hospital level. Torreón Universitario Magazine, 12(33), 103–111. https://doi.org/10.5377/rtu.v12i33.15895
Hall, A. C. (2019). A comparison of DNA stains and staining methods for Agarose Gel Electrophoresis. bioRxiv. https://doi.org/10.1101/568253
Ude, A., Afi-Leslie, K., Okeke, K., & Ogbodo, E. (2022). Trypan Blue Exclusion Assay, Neutral Red, Acridine Orange, and Propidium Iodide. In Cytotoxicity (pp. 1-16). IntechOpen.
Masciotti, V. (2018). Design of an environment-independent, tunable 3D DNA-origami plasmonic sensor. [Source - Unpublished source. Consider removing or including publication details if available.
Debroy, A., Yadav, M., Dhawan, R., Dey, S., & George, N. (2022). DNA dyes: toxicity, remediation strategies, and alternatives. Folia Microbiologica, 67(4), 555-571.
Haines, A. M., Tobe, S. S., Kobus, H. J., & Linacre, A. (2015). Effect of nucleic acid binding dyes on DNA extraction, amplification, and STR typing. Electrophoresis, 36(20), 2561-2568.
Vílchez, G., & Alonso, G. (2009). Scope and limitations of molecular epidemiology methods based on nucleic acid analysis. Journal 1 of the Venezuelan Society of Microbiology, 29(1), 6-12.